Title
Putting Prevention into Action Webinar
Description
Dental experts discuss sealant implementation
Title
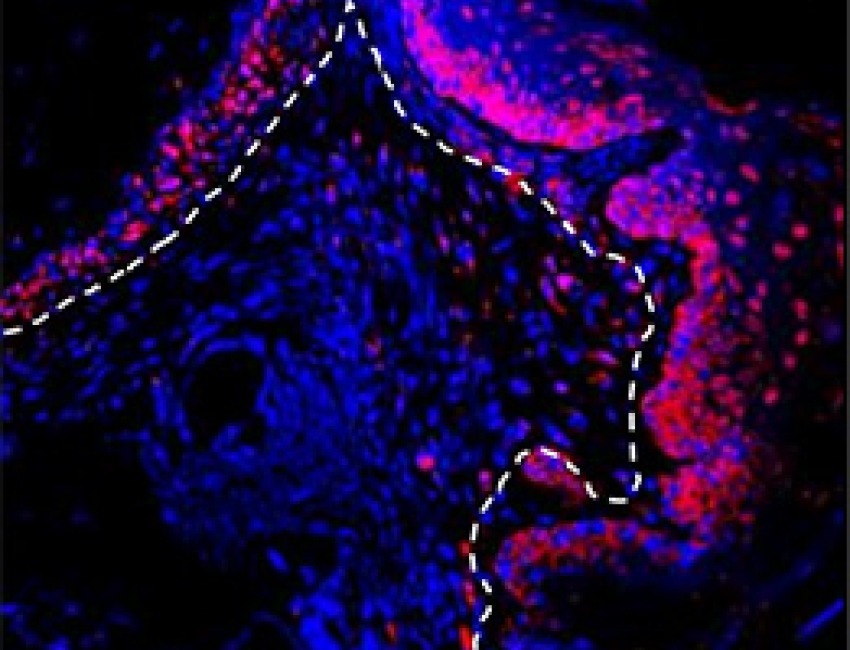
Gum Cells May Trigger Periodontal Disease
Description
Mechanism may recur in other diseases
Title
Congress Honors NIDCR’s Anniversary
Description
Legislation recognizes NIDCR’s role in research and public health
NIDCR News
News/Event Date
News/Event Date
News/Event Date
Research Brief
Upcoming NIDCR Events
Virtual meeting
-
Building 31, Conference Room 6C
Ruth Kirchstein Auditorium, Natcher Conference Center (Bldg. 45), National Institutes of Health (NIH), Bethesda, Maryland
Last Reviewed
April 2024