A novel approach yields genetic insights into how our faces form
You don’t need to know which genes shaped your nose or chin to know which parent to thank (or blame). It’s obvious that our facial traits are highly heritable. Yet, despite scientists’ growing insights into the human genome, surprisingly little is known about facial genetics. Identifying the genes that guide normal development of the face could be key to understanding how that process sometimes goes awry in the case of birth defects and malformations like cleft lip and palate.
To fill in such knowledge gaps, NIDCR in 2009 launched FaceBase, an effort to build a public repository of craniofacial data readily available to all scientists. This centralized database would allow researchers to more efficiently carry out studies to answer critical questions about craniofacial biology.
In an example of how the NIDCR initiative has enabled important scientific advances, a recent study using FaceBase data revealed at least 15 genetic regions strongly linked to differences in face shape in healthy humans. The work emerged from a collaboration among scientists at several universities in the U.S. and Belgium. The findings, published in Nature Genetics, show the power of a new approach that analyzes human faces to identify relevant genes.
The researchers started by examining 3-D facial images and genetic information from a group of more than 2,000 healthy adults of European ancestry. The data had been collected by University of Pittsburgh scientists led by senior author Seth Weinberg, PhD, as part of the first phase of FaceBase in 2009.
To analyze the data, Peter Claes, PhD, of KU Leuven in Belgium, came up with a novel way to parse faces into variables for a widely used method of identifying genes of interest, called a genome-wide association study, or GWAS. For these studies, scientists survey millions of genetic variants across multiple genomes to determine which variants are linked to a disease or trait—facial shape, in this case.
Previous attempts to carry out face-shape GWAS yielded limited information. One reason for this, according to the study authors, is that face-shape measurements in those studies had been predefined in a relatively arbitrary way.
“There’s no rhyme or reason to the traditional ways of measuring the face, so there’s no reason to think those measures have any biological relevance,” says Weinberg. The power of Claes’s approach, he explains, is that “we get rid of those up-front assumptions and allow the data to tell us which traits are likely to be important.”
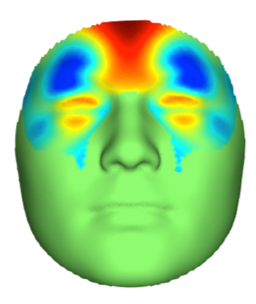
Whereas previous studies used a handful of facial landmarks or simple distances to capture information about facial features, Claes devised mathematical models that employ machine-learning to divide the face into meaningful segments. Claes’s data-driven approach divided the face into 63 segments at differing levels of resolution. For example, the nose region was divided into five segments, each covering increasingly “zoomed in” levels of detail. The researchers then performed a GWAS of each facial segment, enabling a comprehensive look at genetic variations affecting every part of the face.
The results showed that aspects of face shape were significantly linked to gene variants in 38 different regions of the genome. The group repeated the analyses using data from a separate group of 1,719 healthy adults collected by collaborators at Pennsylvania State University led by Mark Shriver, PhD, a senior author on the paper. In this replication study, 15 of the original 38 genomic regions were confirmed to be significantly associated with the same aspects of face shape. At least 11 of these regions had been identified in previous face-shape GWAS, lending further support to the findings. Many of the regions contained genes known to be involved in facial development or implicated in human craniofacial syndromes.
To further explore the biological significance of the GWAS results, the group sought the cell biology expertise of Joanna Wysocka, PhD, and colleagues from Stanford University, who are also generating FaceBase data in a separate project. For the current study, Wysocka and her team examined human neural crest cells, which guide the earliest stages of facial development in the embryo. The scientists reasoned that if variations in human face shape arise in early development, then the 15 genome regions should be active in neural crest cells grown in culture.
This turned out to be the case. Comparing markers of gene activity across 30 types of embryonic and adult cells, the scientists found that activity in the 15 gene regions appeared highest in the neural crest cells, further confirming the GWAS results.
Now, the group is working to extend their analysis to larger and more diverse groups of people, including African and Asian populations. Data on some of these groups currently reside in NIDCR’s FaceBase repository. Increasing the number of people studied will give the researchers higher confidence in the accuracy of their results.
“Studies like ours can provide a roadmap for scientists who are looking to understand how molecules work to shape our faces and the wide range of differences in facial shapes,” says Weinberg.
In addition, the implications of this work can reach far beyond an understanding of normal facial structures. “Identifying genes that underlie shape differences in the faces of healthy populations can help clarify the genetic bases of craniofacial syndromes,” Weinberg says. Such knowledge is critical to finding better ways to prevent and treat these conditions.
Reference
Genome-wide mapping of global-to-local genetic effects on human facial shape. Claes P, Roosenboom J, White JD, Swigut T, Sero D, Li J, Lee MK, Zaidi A, Mattern BC, Liebowitz C, Pearson L, González T, Leslie EJ, Carlson JC, Orlova E, Suetens P, Vandermeulen D, Feingold E, Marazita ML, Shaffer JR, Wysocka J, Shriver MD, Weinberg SM. Nat Genet. 2018 Mar;50(3):414-423. doi: 10.1038/s41588-018-0057-4. Epub 2018 Feb 19. PMID: 29459680.